 Yesterday I went to the annual Thomas S Whittam Memorial Lecture here at MSU. Howard Ochman talked about "Evolutionary Forces Affecting Bacterial Genomes", though he had changed the title to "Determinants of GEnome Size and complexity.
Yesterday I went to the annual Thomas S Whittam Memorial Lecture here at MSU. Howard Ochman talked about "Evolutionary Forces Affecting Bacterial Genomes", though he had changed the title to "Determinants of GEnome Size and complexity.Based on research in his lab had two conclusions about the evolution of bacterial genomes:
- Genome size is drifting
- GC-content is under selection
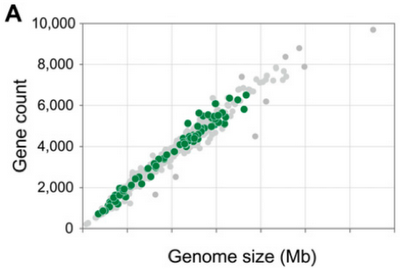
There are trends in the GC-content (amount of guanine and cytosize in the DNA) that differs among bacterial taxa. But closely related species have similar GC-content, and it turns out that genetic drift is not responsible for this, but that it is driven by selection. "Escherichia coli strains harboring G+C-rich versions of genes display higher growth rates" (Raghavan, 2012).
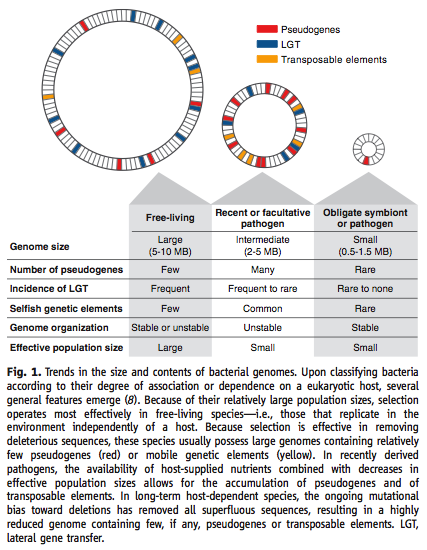
Genome size and abundance of pseudogenes correlates with the size of the effective population size: a larger Ne gives larger genomes, while smaller Ne results in smaller genomes. Pseudogene abundance is less straightforward, with the largest and smallest genomes and Ne both having few pseudogenes, but those intermediate in size having many.
References
Kuo CH, & Ochman H (2010). The extinction dynamics of bacterial pseudogenes. PLoS genetics, 6 (8) PMID: 20700439
Kuo CH, Moran NA, & Ochman H (2009). The consequences of genetic drift for bacterial genome complexity. Genome research, 19 (8), 1450-4 PMID: 19502381
Raghavan R, Kelkar YD, & Ochman H (2012). A selective force favoring increased G+C content in bacterial genes. Proceedings of the National Academy of Sciences of the United States of America, 109 (36), 14504-7 PMID: 22908296
